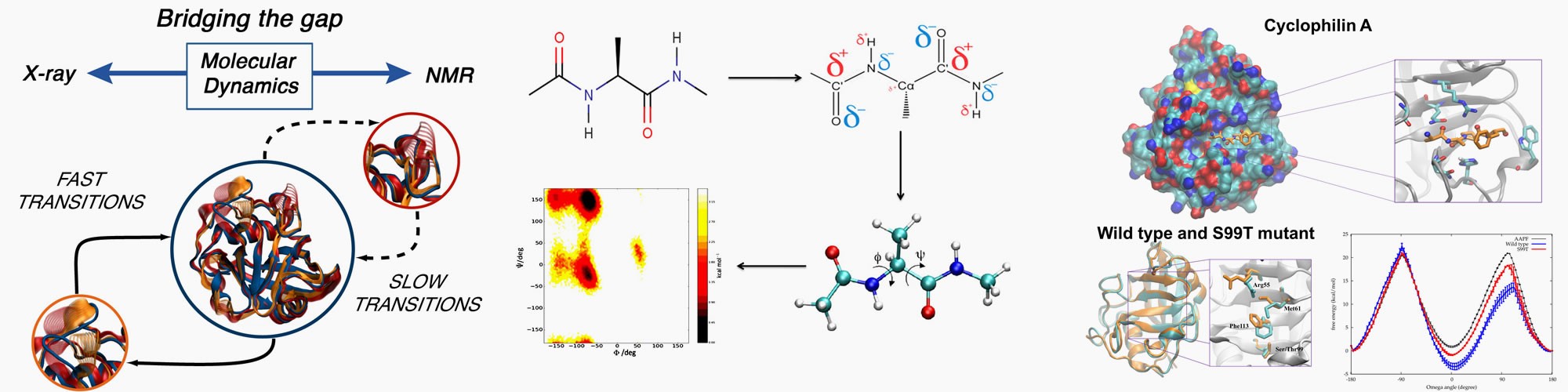
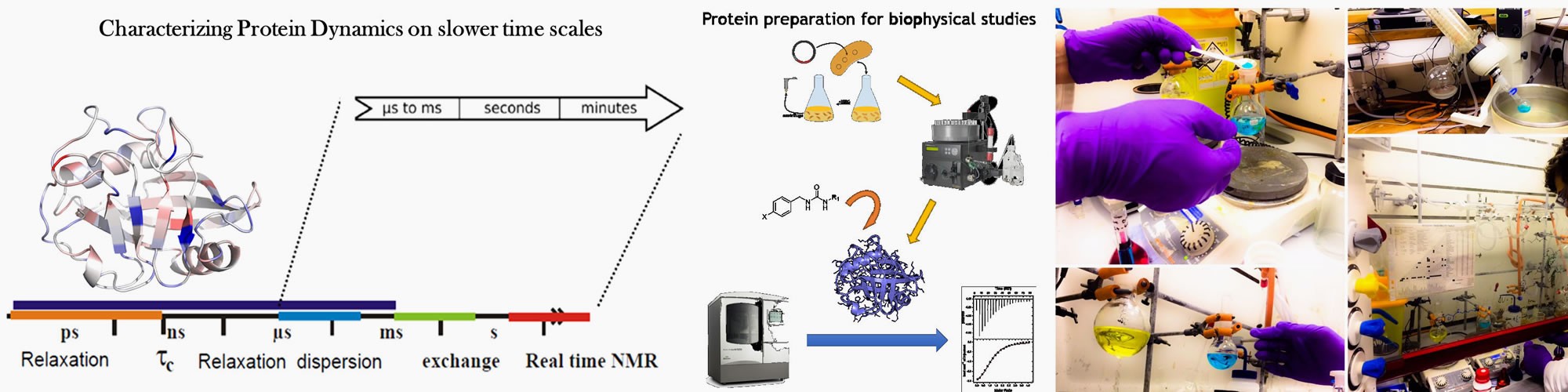
Atomistic description of conformational changes in proteins from pico to millisecond timescales
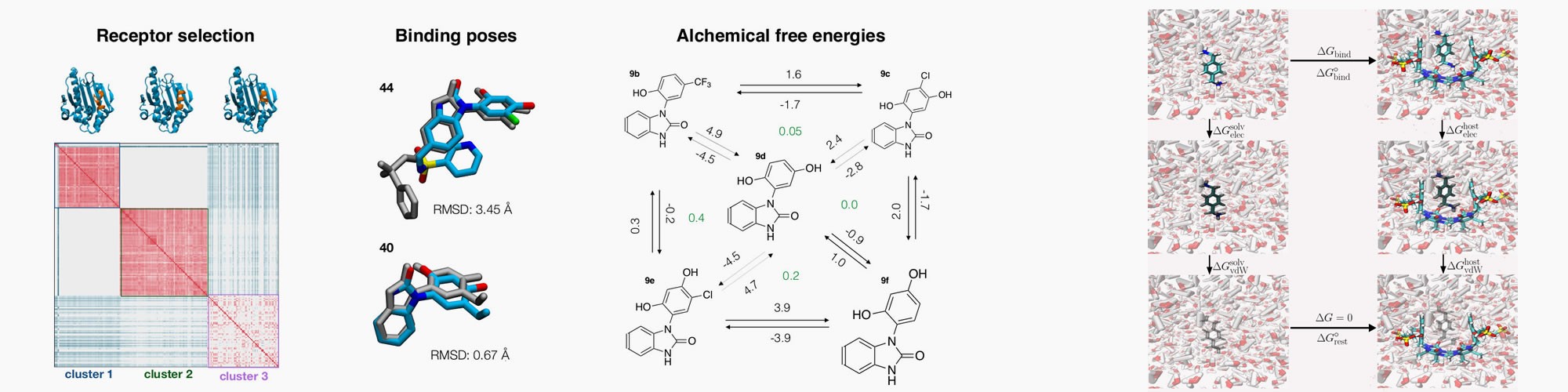
Virtual screens and de-novo ligand design via free energy calculations
Research in the Michel lab is focussed on quantitative description of protein-ligand interactions. This is pursued through development of biomolecular simulation software and algorithms, and biophysical measurements to characterise the structure, dynamics and thermodynamics of protein-ligand interactions.
Currently
PhD studentship opportunity:
''Structure-Based Modelling of Cytochrome P450 Enzyme Inhibition and Metabolism''
Latest News
Group updates
Two group members recently moved on to other pastures. Congratulations to Finlay who passed his PhD viva last month (unfortunately we were all so distracted that we forgot to take...
Wellcome to Silvia
The group is happy to host visiting PhD student Silvia Menin from the University of Padova. Silvia is working in the group of Prof. Stefano Moro in collaboration with...
Young Modellers Forum 2024
Congratulations to Finlay Clark for winning a prize for his talk on ''Automated Adaptive Absolute Binding Free Energy Calculations'' at the 28th Young Modellers Forum meeting in Oxford, November 29th...
New academic year update
The group welcomes new PhD students Benedict Tan and Roy (Haolin) Du. Benedict is working on a molecular glues design project, in collaboration with the Centre for Targeted Protein Degradation...
Meet our group members